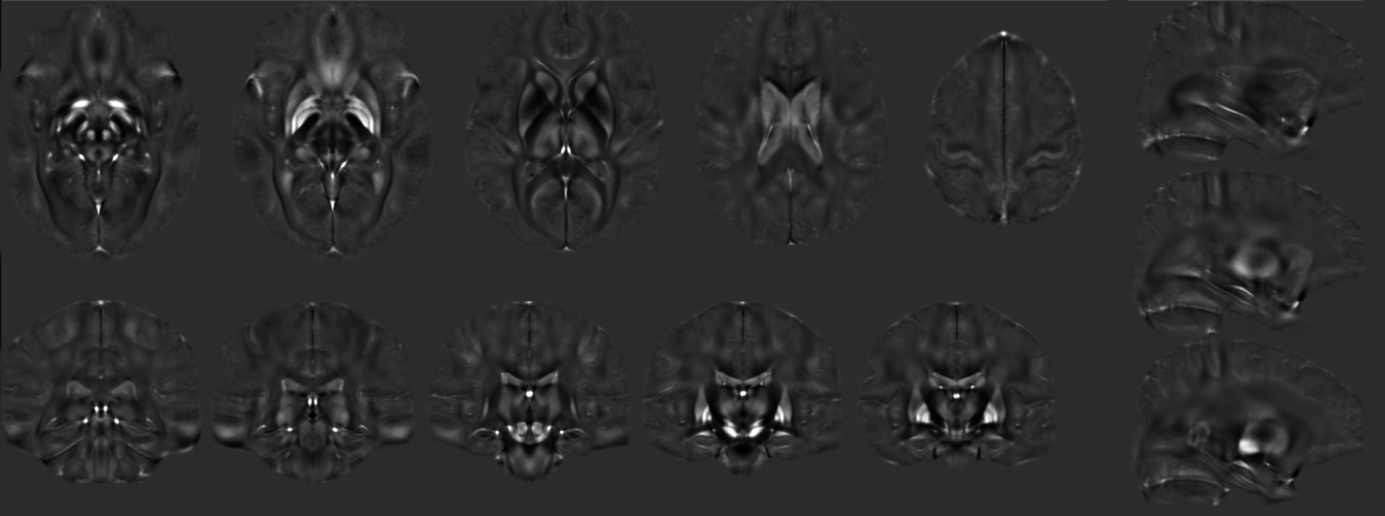
QSM
Downloads
View online and download the model in MINC or NiFTI format here:
GRE model
greModel_L14_gre-7T-weightedMeanOfAllEchoes-asym-mincanon_v0.8 [ View model online | mnc | nii ]
QSM model
qsmModel_L14_gre-7T-weightedMeanOfAllEchoes-asym-mincanon_v0.8 [ View model online | mnc | nii ]
Citing this model:
Bollmann, Steffen, Andrew Janke, Lars Marstaller, David Reutens, Kieran O’Brien, and Markus Barth. “GRE and QSM Average 7T Model” January 1, 2017. doi:10.14264/uql.2017.178.
References describing this model:
Bollmann, Steffen, Simon Daniel Robinson, Kieran O’Brien, Viktor Vegh, Andrew Janke, Lars Marstaller, David Reutens, and Markus Barth. “The Challenge of Bias-Free Coil Combination for Quantitative Susceptibility Mapping at Ultra-High Field.” Magnetic Resonance in Medicine, March 1, 2017, n/a-n/a. doi:10.1002/mrm.26644.
Janke AL, O’Brien K, Bollmann S, Kober T, Marstaller L, Barth M. A 7T Human Brain Microstructure Atlas by Minimum Deformation Averaging at 300um. In 24th Annual ISMRM Scientific Meeting and Exhibition, Singapore.
Acquisition details
QSM: 29 (14 female, 26.6±3.8yr) individuals were imaged using a multiple echo gradient recalled echo 3D whole brain dataset. TR=25ms, TE=4.4,7.25,10.2,13.25,16.4, 19.65,23ms flip angle=13, FOV 210x181.5x120mm, matrix=280x242x160, GRAPPA=2. The phase data was combined using COMPOSER [8] and susceptibility processing was performed using total generalized variation (TGV) [9], which incorporates phase unwrapping, background field removal and dipole inversion in a single step.
- Robinson SD, Wolfgang Bogner, Barbara Dymerska, Pedro Cardoso, Günther Grabner, Xeni Deligianni, et al. COMbining Phased array data using Offsets from a Short Echo-time Reference scan (COMPOSER). ISMRM 23rd Annual Meeting & Exhibition. 2015
- Langkammer, C., Bredies, K., Poser, B.A., Barth, M., Reishofer, G., Fan, A.P., Bilgic, B., Fazekas, F., Mainero, C., Ropele, S., 2015. Fast quantitative susceptibility mapping using 3D EPI and total generalized variation. NeuroImage 111, 622–630. doi:10.1016/j.neuroimage.2015.02.041
Preview